From cells to purified capsids: How to develop a scalable rAAV process
Cell & Gene Therapy Insights 2022; 8(3), 611–619
DOI: 10.18609/cgti.2022.094
Adeno-associated viral (AAV) vectors are widely used for gene therapy, with multiple serotypes and several different synthetic capsid variants targeting different tissues. The number of AAVs in clinical trials has increased over recent years and the serotypes primarily used have changed, from AAV1 and AAV2 initially to AAV8 and AAV9 today.
Within our AAV process at Cytiva, on the upstream side, we use triple plasmid transfection and HEK293 cells in suspension and AAV expressing green fluorescent protein (GFP) as a model system. We use scalable, single-use bioreactors. In the downstream process, steps include cell lysis and DNA fragmentation, clarification, concentration and buffer exchange, capture, polishing, formulation and finally, sterile filtration. We have a large analytical package including various assays for infectious virus titer, total virus titer, and viral genome (VG) titer. We also utilize different assays to measure full/empty ratio of samples and host cell impurities, and for vector characterization.
This article describes the production and purification process of several AAV serotypes and the optimization of each process step, with a particular focus on ensuring high overall yields of full capsids, empty capsid reduction, and efficient impurity removal. We will explore common pitfalls for rAAV processing and ways to overcome the challenges presented. We will also discuss how full/empty AAV capsid separation for AAV2, AAV5, AAV8, and AAV9 serotypes can be significantly
improved with a single chromatography resin and one protocol.
Upstream cell culture & virus production
Transient transfection optimization
We used a design of experiment (DoE) approach to optimize and develop a new transfection protocol for production of recombinant AAV5 (rAAV5). We confirmed that DNA concentration affects vector genome (VG) titer and percentage full capsids. As DNA concentration increases, the VG titer also increases, whilst the percentage full capsids decreases. This poses some difficulty, because it is desirable to have both high VG titer and high percentage full capsids. In order to balance these variables, we used 0.75% μg/mL of DNA. We also discovered that a short incubation time of the transfection mix prior to application to the suspension cell culture significantly increased the % full capsids.
Based on our DoE results, we developed a workflow for rAAV5 production. The transfection parameters we selected included a viable cell density of 1×106 cells/mL, a polyethylenimine (PEI) transfection reagent in a 2:1 ratio with DNA, a transfection volume of 5% of total volume, and an incubation of 15 min at room temperature with a DNA plasmid ratio of 1:1:2 (rep/cap:helper:transgene GFP).
AAV production in a single-use bioreactor
We performed three 10 L rAAV5 production runs in a stirred tank bioreactor, which were reproducible both with VG and viral particle (VP) titers. The percentage full capsids were all above 10%.
As can be seen in Figure 1, this AAV production process is scalable from 20 mL to 25 L and is consistent across various AAV serotypes. We reached approximately 1014 VP/L and 1013 VG/L. The range of percentage full capsids was 10–40%.
Downstream purification process
Harvest & filtration
To develop the purification process for recombinant AAV5, we firstly considered the harvest and filtration steps. The harvest was completed by cell lysis, DNA fragmentation and clarification, using Tween™ 20, followed by nuclease treatment directly in the bioreactor. Filtration was performed with three different filter capsules. We achieved recovery of up to 80% over the harvest and clarification steps. Concentration and buffer exchange was completed by tangential flow filtration using hollow fibers. We achieved efficient impurity removal with low loss of virus using 300 KDa molecular weight cut-off (MWCO).
We performed affinity capture chromatography with the Capto™ AVB, HiTrap™ column. The elution was different for AAV2 and AAV5, with AAV5 recovery being negatively affected by salt in the elution buffer, and we used a lower elution pH compared to AAV2. However, using the respectively optimized protocols the performance was similar, with a recovery of up to 90%. The binding capacity was 2–5×1014 VP/mL with a 100-fold or higher concentration. Even without salt in the eluate, we confirmed with analytical
sequencing that aggregation was below 1%.
Full/empty capsid separation
The resultant highly concentrated, pure AAV sample, containing both full and empty capsids, required a polishing step using ion exchange chromatography to reduce the empty capsids. Full capsids have a lower pI than empty capsids (5.9 versus 6.3 on average), and this charge difference can be used in anion or cation exchange with salt elution to separate the capsids. In our experience, anion exchange is preferable for this process. At higher pH, the net charge is negative, and the less-charged empty capsids will elute first in a salt gradient.
During anion exchange to reduce empty capsids for AAV5, we discovered that the MgCl2 concentration was critical for the separation. We used bis-tris propane buffer at pH 9 to compare the separation results using 0, 5, 17, or 20 mM MgCl2 at a constant concentration, and then applied a linear NaCl gradient. We identified that high concentrations of 15–20 mM of MgCl2 were optimal to maximize separation.
The final protocol we used for scaling up the AAV5 polishing step with Capto Q ImpRes resin to 51 mL HiScale™ column is shown in Figure 2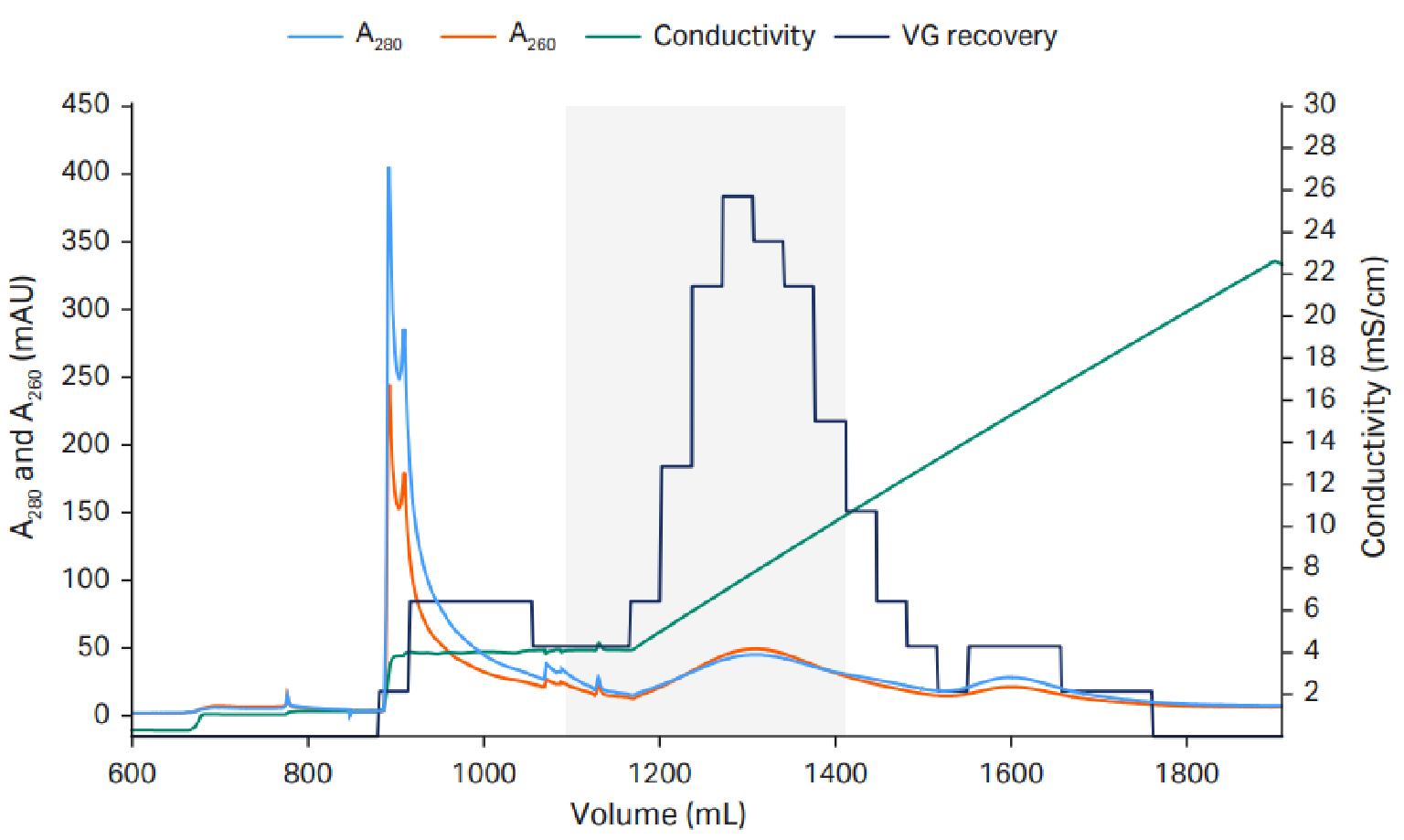 Anion exchange to reduce empty capsids.AAV final protocol and scale up to 10 L.. We used 20 mM tris pH 9 with a constant 18 mM MgCl2 concentration and a linear NaCl gradient up to 200 mM.
Anion exchange to reduce empty capsids.AAV final protocol and scale up to 10 L.. We used 20 mM tris pH 9 with a constant 18 mM MgCl2 concentration and a linear NaCl gradient up to 200 mM.
The empty capsids eluted in an elution hold step with 18 mM MgCl2 before the linear NaCl gradient, in which the full capsids eluted.. VG recovery was measured using quantitative PCR (qPCR). The capacity was approximately 1–3×1013 VP/mL resin.
We achieved approximately 60–70% VG recovery and 40–65% full capsids. The result varies depending on the fractions that are pooled, as there is a tradeoff between VG recovery and percentage full capsids.
From our three repeated 10 L AAV5 process runs, we saw virus titer results of approximately 1015 VP/L in the final samples. Percentage full capsids was determined in three different ways: qPCR/ELISA, qPCR/surface plasmon resonance (SPR), and anion exchange chromatography (AIEX) high-performance liquid chromatography (HPLC). The result varied depending on the method used, ranging from 42–71%. Regulatory purity targets were met, as the host cell protein (ELISA), host cell DNA (qPCR) were below 100 ng and 10 ng per dose (1011–1012 VG) respectively, and residuals levels (nuclease and Tween 20) were below the limit of detection.
Improving the AAV polishing step
Dextran extenders and MgCl2
We further explored the AAV polishing step to attempt to improve the percentage full capsids and VG recovery.
We found that dextran extenders enhance full/empty capsid separation (Figure 3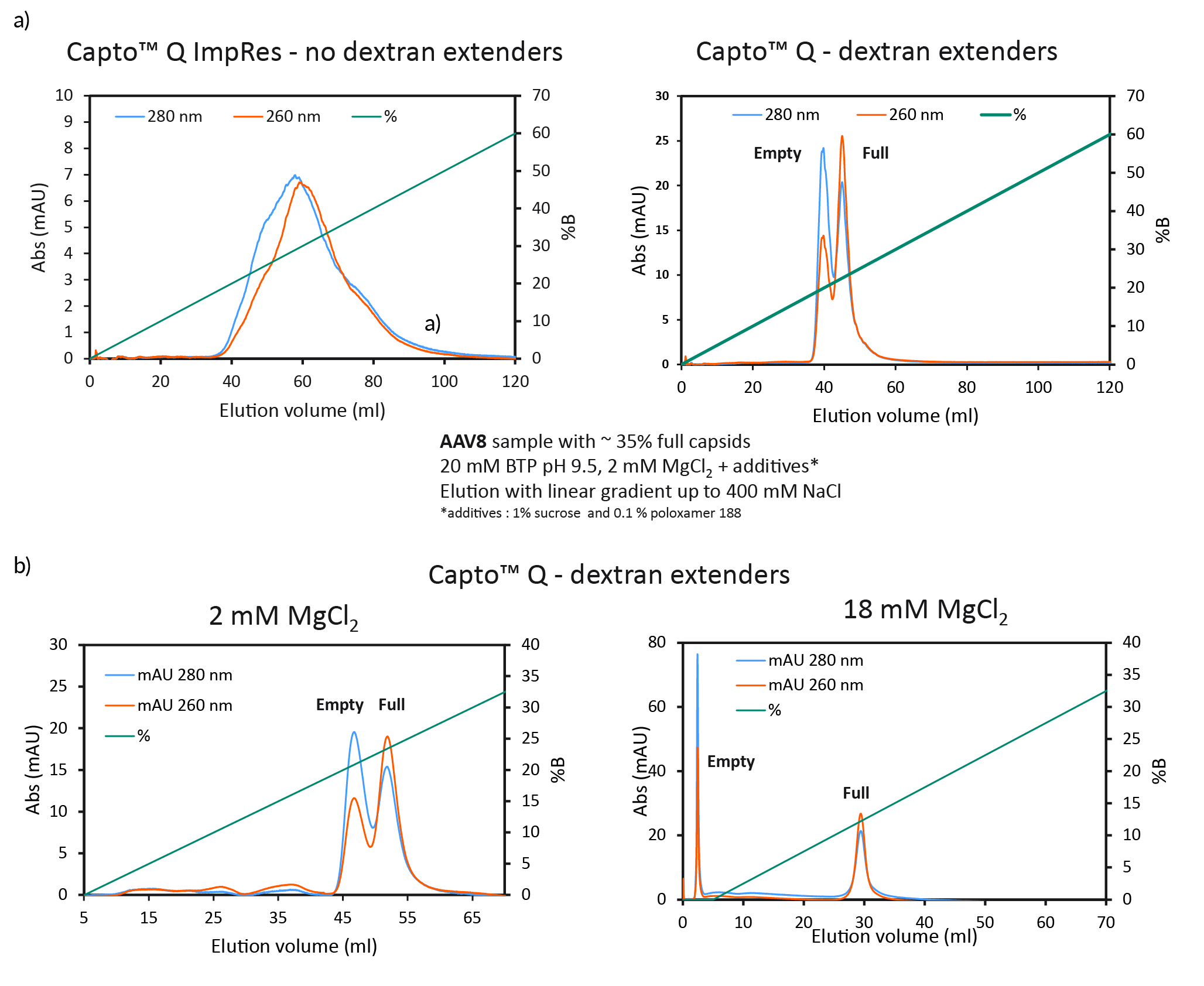 Full/empty capsid separation with Capto Q using isocratic step elution.a) Full/empty capsid separation, without (Capto Q ImpRes) and with (Capto Q) dextran extenders at 2mM MgCl2. b) Full/empty capsid separation with Capto Q, with dextran extenders, at 2 or 18 mM MgCl2.a). Without dextran extenders, the peaks overlapped. When present on the resin, the dextran extenders enhanced the separation of the peaks. We also found that higher constant concentration of MgCl2 also enhanced the separation and resulted in earlier elution and base line separation (Figure 3b).
Full/empty capsid separation with Capto Q using isocratic step elution.a) Full/empty capsid separation, without (Capto Q ImpRes) and with (Capto Q) dextran extenders at 2mM MgCl2. b) Full/empty capsid separation with Capto Q, with dextran extenders, at 2 or 18 mM MgCl2.a). Without dextran extenders, the peaks overlapped. When present on the resin, the dextran extenders enhanced the separation of the peaks. We also found that higher constant concentration of MgCl2 also enhanced the separation and resulted in earlier elution and base line separation (Figure 3b).
Next, we wanted to investigate isocratic step elution with NaCl at high constant concentration of MgCl2 (Figure 4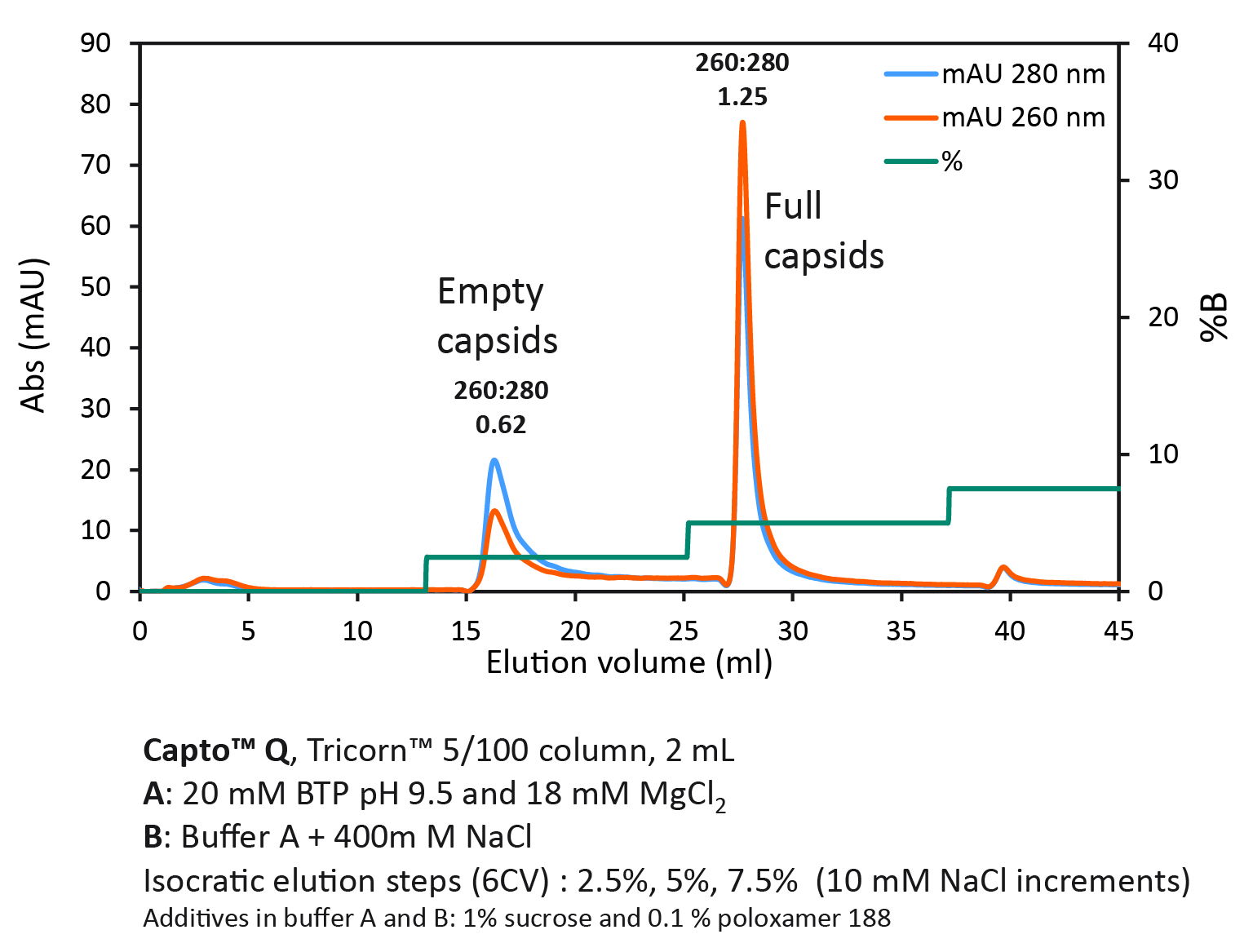 Full/empty capsid separation with Capto Q using isocratic step elution.Using a step gradient, we achieved a UV ratio 260:280 of 1.25 in the second full capsid peak, indicating a high purity of full capsids.).
Full/empty capsid separation with Capto Q using isocratic step elution.Using a step gradient, we achieved a UV ratio 260:280 of 1.25 in the second full capsid peak, indicating a high purity of full capsids.).
Multi-serotype protocol for polishing with Capto Q
Capto Q (with dextran extenders) equilibrated with 20 mM Na-acetate, pH 9 containing 2 mM MgCl2 (buffer A) and after sample application (up to approx. 1013VP/ml resin and conductivity 1–3 mS/cm) a buffer B containing 250 mM Na-acetate was used to elute empty and full capsids respectively. To identify the optimal elution conditions for all serotypes studied (AAV2, AAV5, AAV8, and AAV9), a pre-screening procedure with 5% B buffer incremental steps was performed (Figure 5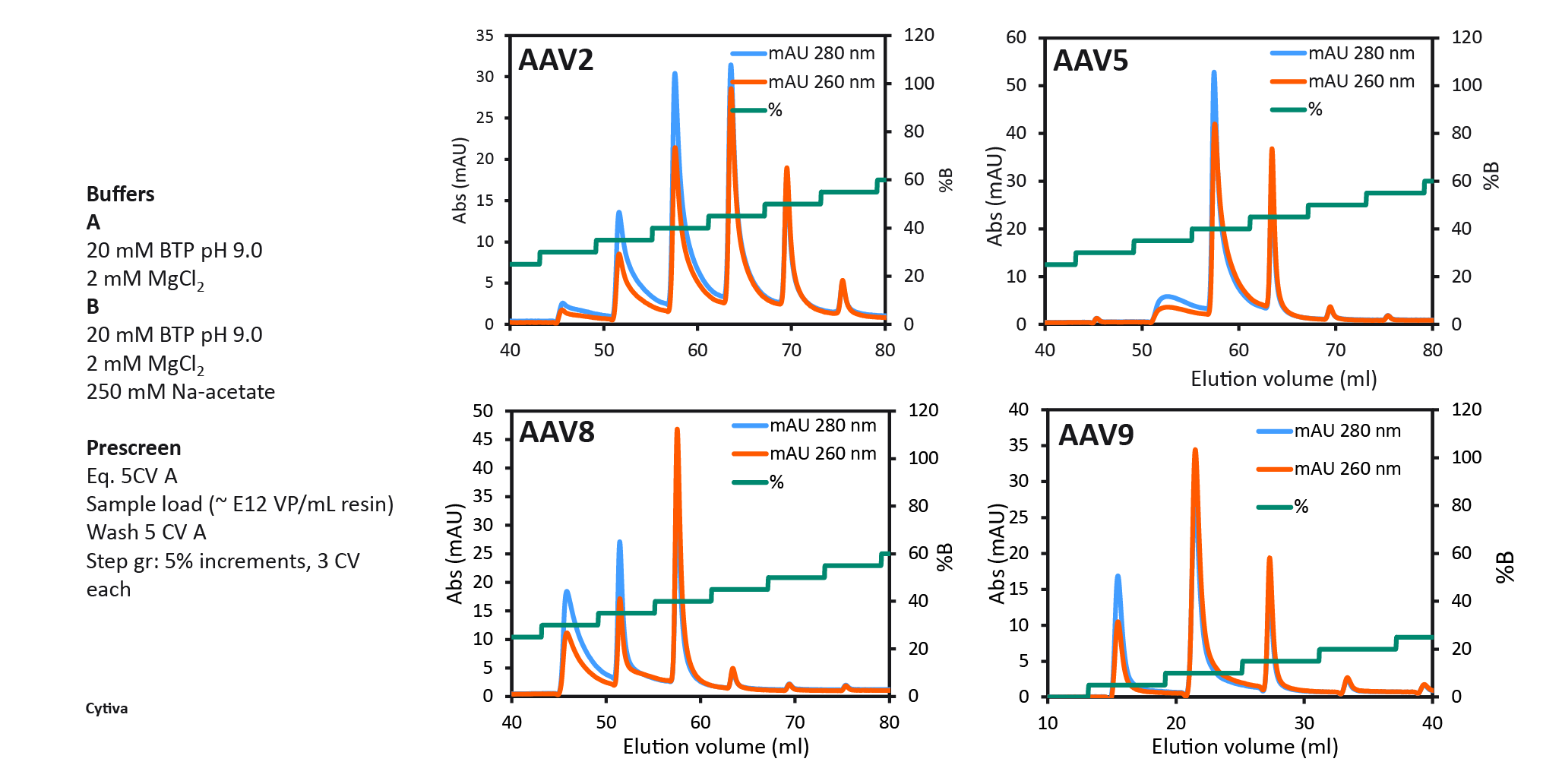 Capto Q pre-screening to select two elution step conditions. 100% B buffer contained 250 mM sodium acetate as elution salt.). This enabled the selection of the %B for the first elution step in the final 2 step protocol. The % B buffer at which the first empty peak eluted was selected for elution step 1, eluting the empty capsids, followed by the second elution step with up to 100% B buffer to elute the full capsids.
Capto Q pre-screening to select two elution step conditions. 100% B buffer contained 250 mM sodium acetate as elution salt.). This enabled the selection of the %B for the first elution step in the final 2 step protocol. The % B buffer at which the first empty peak eluted was selected for elution step 1, eluting the empty capsids, followed by the second elution step with up to 100% B buffer to elute the full capsids.
The final protocol that was selected for all serotypes is shown in Figure 6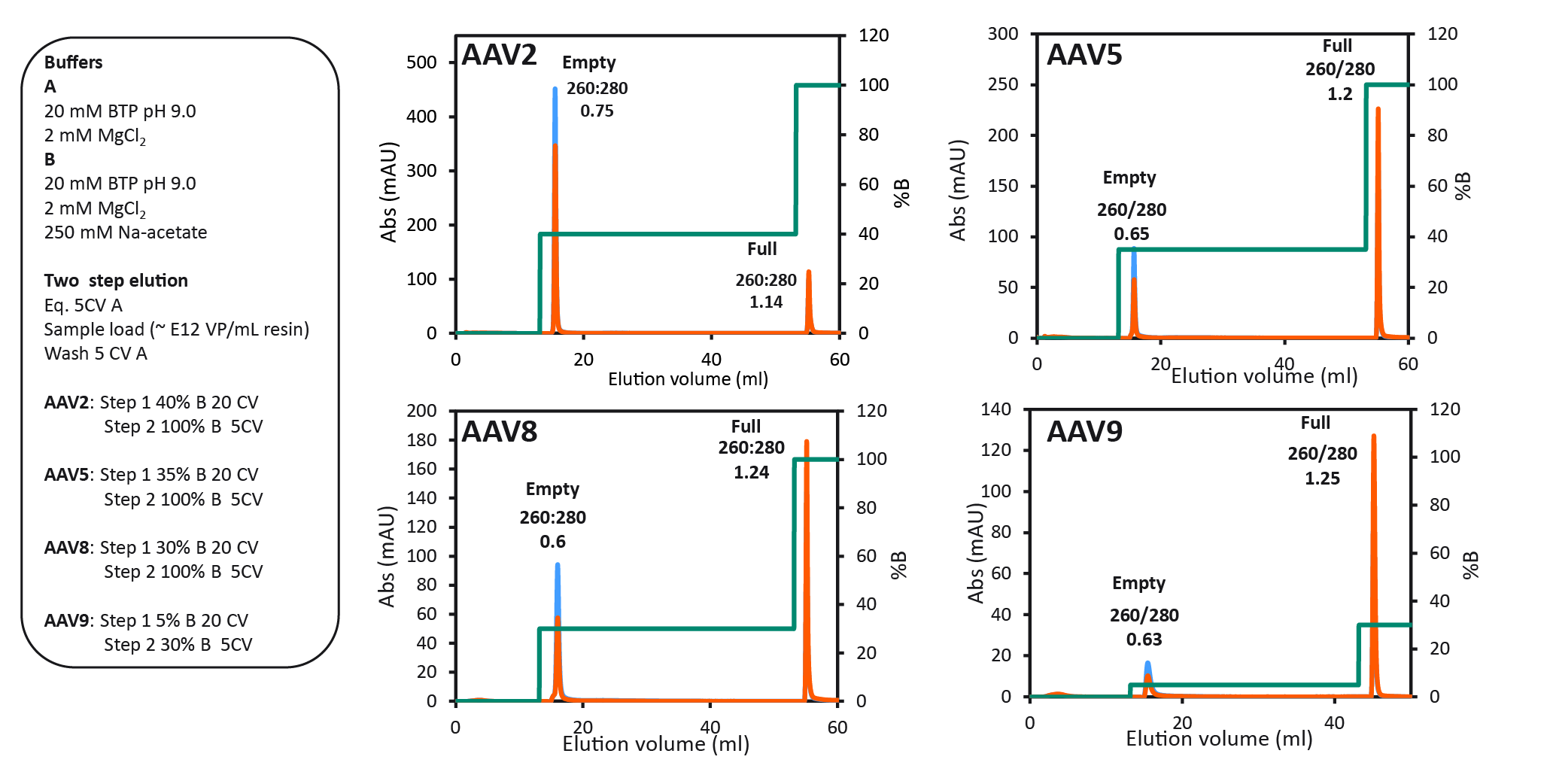 The final two-step protocol with baseline separation of full and empty for four different AAV serotypes.. Between all four serotypes, we saw variation of the optimal %B buffer for step one. The data show the binding strength to the anion exchange resin as follows: AAV2 > AAV5 > AAV8 > AAV9. AAV9 binding was the weakest, seen by the decreasing % B buffer required to elute the empty capsids. 40% B buffer was required for eluting AAV2 empty capsids and only 5% B buffer was needed to elute AAV9 empty capsids.
The final two-step protocol with baseline separation of full and empty for four different AAV serotypes.. Between all four serotypes, we saw variation of the optimal %B buffer for step one. The data show the binding strength to the anion exchange resin as follows: AAV2 > AAV5 > AAV8 > AAV9. AAV9 binding was the weakest, seen by the decreasing % B buffer required to elute the empty capsids. 40% B buffer was required for eluting AAV2 empty capsids and only 5% B buffer was needed to elute AAV9 empty capsids.
Step one is prolonged over 20 column volumes, to maximize the elution of empty capsids before the elution of full capsids. This protocol can be used for most capsid types including engineered variants, if the pre-screening procedure is firstly used to determine the exact %B buffer needed for the step 1 eluting the empty capsids. The UV ratios indicating the full to empty capsid ratio have also been confirmed with qPCR and ELISA. The results are summarized
in Table 1.
| Table 1 Summary of final two-step protocol results with each AAV serotype. | |||||||
| Serotype | Start sample | Peak 1 (empty capsids) | Peak 2 (full capsids) | ||||
| qPCR:ELISA (% full capsids) | UV 260:280 (peak area) | VG recovery (%) | qPCR:ELISA (% full capsids) | UV 260:280 (peak area) | VG recovery (%) | qPCR:ELISA (% full capsids) | |
| AAV2 | 7–10% | 0.75 | NA | NA | 1.14 | NA | NA |
| AAV5 | 47% | 0.65 | 7 | 5 | 1.20 | 80 | 100 |
| AAV8 | 11–35% | 0.60 | 3 | 1 | 1.24 | 80 | 95 |
| AAV9 | 40% | 0.63 | 0.3 | 1 | 1.25 | 91 | 100 |
Analytics
Analytics are critical for a successful AAV process development and manufacturing but time consuming. Knowledge about the detection range is needed to know the compatibilities with samples of different concentration. Assay variation, including inter- and intra-assay reproducibility, is required in order to know if and how small differences in the result may be significant and/or reliable. Controls and references should be included in every analysis run to determine accuracy, and any assay inhibiting or enhancing effect from the process buffers must be checked to minimize matrix or sample buffer interference. Orthogonal methods for full and empty capsid analysis are also critical, as qPCR and ELISA ratios can be variable. Finally, any simplification and/or automation where possible will reduce throughput time. We recommend spending time for assay validation as it will facilitate process optimization and bring confidence in the process quality and productivity.
We have developed antibody-based Biacore SPR assays for AAV2 and AAV5, which are compatible for AAV samples from cell culture to final product. The assays are simple and easy to use, with improved precision and a higher degree of automation compared to ELISA.
Summary
We have a scalable HEK293 suspension cell culture AAV production process, with animal-derived component-free medium in single-use bioreactors. Our triple plasmid transfection protocol shows high productivity of viral genomes and high percentage full capsids, and is suitable for use with AAV2, AAV5, AAV8 and AAV9 serotypes. We have a scalable, start-to-finish AAV5 production process and a polishing protocol with anion exchange using Capto Q. Orthogonal methods determining percentage full capsids are needed and we have developed reproducible, easy-to-use Biacore assays for AAV2 and AAV5. This start-to-finish AAV process is suitable for large-scale, clinical-grade production.
Q&A with Åsa Hagner McWhirter
How closely do the empty/full ratios via qPCR or capsid ELISA correlate with the area under the curve (AUC), and other analytical methods for determining empty/full ratios?
AHM: It is important to use more than one assay, because qPCR and ELISA assays can vary. A ratio between them can give uncertain values, but you cannot use analytical ultracentrifugation or cryo TEM on all your samples during process development due to high cost and long time to results. You need to do some selection of samples for those high-end analysis. There are many different methods you can use to more quickly confirm those qPCR and ELISA results, such as analytical anion exchange or size exclusion combined with UV 260:280 detection for example.
How do you determine infectious titer by flow cytometry?
AHM: This can be done using a cell-based assay that measures transduction.
The insert here is GFP. When the cells are transduced, GFP will be expressed. You incubate different dilutions of your virus sample with the cells, detach the cells, and then run them in flow cytometry. This allows you to quantify the GFP expression as a measurement of the activity of the virus and give you a functional titer.
Was the Capto Q protocol tested with any other AAV serotype?
AHM: Yes - we have collaborated with customers to try out this protocol, and it looks promising. We definitely think that it will be possible for other serotypes, including AAV6 and engineered capsids. However, right now, the data is preliminary and has not yet been published.
Affiliation
Åsa Hagner McWhirter, PhD
Principal Scientist
Cytiva
Authorship & Conflict of Interest
Contributions: All named authors take responsibility for the integrity of the work as a whole, and have given their approval for this version to be published.
Acknowledgements: None.
Disclosure and potential conflicts of interest: The author declare a licence with Sanofi and 2 patents pending. They have no other conflicts of interest.
Funding declaration: The authors received no financial support for the research, authorship and/or publication of this article.
Article & copyright information
Copyright: Published by Cell and Gene Therapy Insights under Creative Commons License Deed CC BY NC ND 4.0 which allows anyone to copy, distribute, and transmit the article provided it is properly attributed in the manner specified below. No commercial use without permission.
Attribution: Copyright © 2022 Cytiva. Published by Cell and Gene Therapy Insights under Creative Commons License Deed CC BY NC ND 4.0.
Article source: This article has been written based on a webinar, which can be found here.
Submitted: May 03 2022; Revised manuscript received: Jun 2 2022; Publication date: Jun 21 2022.



